- Bertholomey ML, Stone K, Lam TT, Bang S, Wu W, Nairn AC, Taylor JR, Torregrossa MM*. Phosphoproteomic Analysis of the Amygdala Response to Adolescent Glucocorticoid Exposure Reveals G-Protein Coupled Receptor Kinase 2 as a Target for Reducing Motivation for Alcohol. Proteomes. 2018 Oct 12;6(4). pii: E41. doi: 10.3390/proteomes6040041. PMID:30322021 [Publication with FRP.]
- Anthonymuthu T, Kenny E, Shrivastava I, Tyurina YY, Hier Z, Ting H-C, Dar H, Tyurin V, Nesterova A, Amoscato A, Mikulska-Ruminska K, Rosenbaum J, Mao G, Jinming Z, Conrad M, Kellum J, Wenzel S, VanDemark A, Bahar I, Kagan V, Bayir H (2018) Empowerment of 15-lipoxygenase catalytic competence in selective oxidation of membrane ETE-PE to ferroptotic death signals, HpETE-PE. J Am Chem Soc 2018, 140 (51), pp 17835-17839 PMID: 30525572.
- Wang K, Steer E, Otero PA, Bateman N, Cheng MH, Scott A, Wu C, Bahar I, Shih Y-T, Hsueh Y-P, Chu C (2018) PINK1 Interacts with VCP/p97 and Activates PKA to Promote NSFL1C/p47 Phosphorylation and Dendritic Arborization in Neurons. eNeuro 5 (6) ENEURO.0466-18.2018. PMID: 30783609
- Wu W*#, Bang S#, Bleecker ER, Castro M, Denlinger L, Erzurum SC, Fahy JV, Fitzpatrick AM, Gaston BM, Hastie AT, Israel E, Jarjour NN, Levy BD, Mauger DT, Meyers DA, Moore WC, Peters M, Phillips BR, Phipatanakul W, Sorkness RL, Wenzel SE* (# co-first authors). Multiview cluster analysis identifies variable corticosteroid response phenotypes in severe asthma”, Am J Resp Crit Care Med (accepted) [Publication with FRP.]
- Zhou L*, Zhou S, Yang P, Tian Y, Feng Z, Xie XQ*, Liu Y*. Targeted inhibition of the type 2 cannabinoid receptor is a novel approach to reduce renal fibrosis. Kidney Int. 2018 Oct;94(4):756-772. doi: 10.1016/j.kint.2018.05.023. Epub 2018 Aug 6. PMID:30093080.
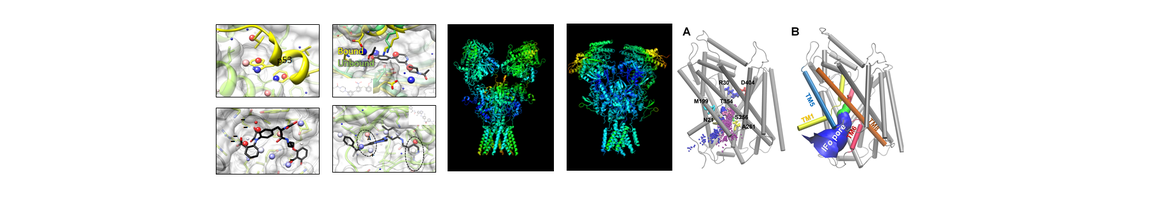
- Lee JY, Krieger J, Herguedas B, García-Nafría J, Dutta A, Shaikh SA, Greger IH*, Bahar I*. Druggability simulations and X-ray crystallography reveal a ligand-binding site in the GluA3 AMPA receptor N-terminal domain. Structure. 2018 Nov 3. pii: S0969-2126(18)30378-2. doi: 10.1016/j.str.2018.10.017. [Epub ahead of print] PMID: 30528594
- Wang H, Lengerich BJ, Aragam B, Xing EP*. Precision Lasso: Accounting for Correlations and Linear Dependencies in High-Dimensional Genomic Data. Bioinformatics. 2018 Sep 1. doi: 10.1093/bioinformatics/bty750. PMID:30184048
- van Dijk L, Giladi M, Refaeli B, Hiller R, Cheng MH, Bahar I*, Khananshvili D*. Key residues controlling bidirectional ion movements in Na+/Ca2+ exchanger. Cell Calcium. 2018 Dec;76:10-22. PMID:30248574
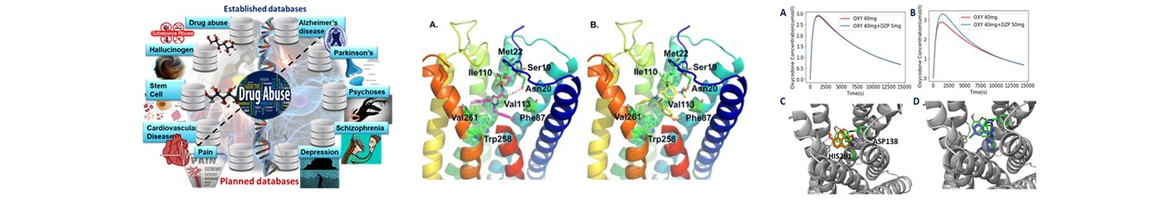
- Wu X, Xie S, Wang L, Fan P, Ge S, Xie XQ, Wu W*. A computational strategy for finding novel targets and therapeutic compounds for opioid dependence. PLoS One. 2018 Nov 7;13(11):e0207027. doi: 10.1371/journal.pone.0207027. eCollection 2018. PMID:30403753 [Joint publication for Cores A and C.]
- Hu Z, Wang L, Ma S, Kirisci L, Feng Z, Xue Y, Klunk WE, Kamboh MI, Sweet RA, Becker J, Lv Q, Lopez OL*, Xie XQ*. Synergism of antihypertensives and cholinesterase inhibitors in Alzheimer’s disease. Alzheimers Dement (NY). 2018 Oct 14;4:542-555. PMID:30386819
- Wang, J; Ge, Y; Xie X-Q. Development and testing of druglike screening libraries. J. Chem. Inf. Model., Article ASAP (DOI: 10.1021/acs.jcim.8b00537).
- Man, V; He, X; Derreumaux, P; Ji, B; Xie X-Q; Nguyen, P; Wang, J. Effects of all-atom molecular mechanics force fields on Amyloid peptide assembly: the case of Aβ16-22 Dimer. J. Chem. Theor. Comput. (accepted) (Manuscript ID ct-2018-01107f.R1).
- Wang, J; Cieplak, P; Luo, R; Duan, Y. Development of Polarizable Gaussian Model for Molecular Mechanical Calculations I: Atomic Polarizability Parameterization to Reproduce Ab Initio Anisotropy. J. Chem. Theor. Comput. (accepted) (Manuscript ID ct-2018-006033.R2)
- He, X.; Man, V.; Ji, B.; Xie, X-Q; Wang, J. Calculate protein-ligand binding affinities with the extended linear interaction energy method: application on the Cathepsin S set in the D3R Grand Challenge 3. J. Comput.-Aided Mol. Des. 2018, Article ASAP (DOI: 10.1007/s10822-018-0162-6)
- Wang K, Steer E, Otero PA, Bateman N, Cheng MH, Scott A, Wu C, Bahar I, Shih Y-T, Hsueh Y-P, Chu CT* (2018) PINK1 interacts with VCP/p97 and activates PKA to promote NSFL1C/p47 phosphorylation and dendritic arborization in neurons. eNeuro, in press. PMID: pending
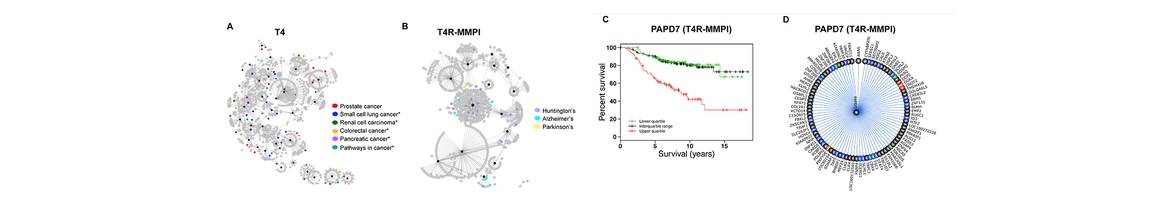
- Pei F, Li H, Liu B*, and Bahar I*. Quantitative systems pharmacological analysis of drugs of abuse reveals the pleiotropy of their targets and the effector role of mTORC1. Frontiers in Pharmacology, accepted with minor revision, doi: 10.1101/470922. PMID: pending
- Dominik, D.; Man, V.; Van-Oanh, N.-T.; Wang, J.; Kawasaki, T.; Derreumaux, P.; Nguyen, P. Breaking down cellulose fibrils with a mid-infrared laser, Cellulose, 2018, 25, 5553-5568.
- Xue Y, Feng ZW, Li XY, Hu ZH, Xu Q, Wang Z, Cheng JH, Shi HT, Wang QB, Wu HY, Xie X.-Q*. The efficacy and safety of cilostazol as an alternative to aspirin in Chinese patients with aspirin intolerance after coronary stent implantation: a combined clinical study and computational system pharmacology analysis. Acta Pharmacologica Sinica. 2018 Feb;39(2):205. PMID: 28933424 PMCID: PMC5800472
- Cha-Molstad H, Lee SH, Kim JG, Sung KW, Hwang J, Shim SM, Ganipisetti S, McGuire T, Mook-Jung I, Ciechanover A, Xie XQ, Kim BY, Kwon YT. Regulation of autophagic proteolysis by the N-recognin SQSTM1/p62 of the N-end rule pathway. 2018 Jan 29:1-3. doi: 10.1080/15548627.2017.1415190. [Epub ahead of print] PMID:29261001
- Yoo YD, Mun SR, Ji CH, Sung KW, Kang KY, Heo AJ, Lee SH, An JY, Hwang J, Xie X-Q, Ciechanover A, Kim BY, Kwon YT. N-Terminal Arginylation Generates a Bimodal Degron That Modulates Autophagic Proteolysis. Proc Natl Acad Sci USA. 2018:201719110; PMCID: PMC5866579.
- Bian Y, Xie X-Q. Computational Fragment-Based Drug Design: Current Trends, Strategies, and Applications. AAPS J. 2018;20(3):59. PMID: 29633051 PMCID: PMC6618289 DOI: 10.1208/s12248-018-0216-7.
- Jing Y, Bian Y, Hu Z, Wang L, Xie X-Q. Deep Learning for Drug Design: An Artificial Intelligence Paradigm for Drug Discovery in the Big Data Era. AAPS J. 2018;20(3):58. PMID: 29603063 PMCID: PMC6608578
- Lengerich BJ, Aragam B, Xing EP. Personalized Regression Enables Sample-Specific Pan-Cancer Analysis. Bioinformatics (ISMB 2018). 2018;34(13):i178-86. doi: 10.1093/bioinformatics/bty250. PubMed PMID: 29949997; PMCID: PMC6022603.
- Sachan M, Xing EP. Self Training for Jointly Learning to Ask and Answer Questions. Proceedings of The 16th Annual Conference of the North American Chapter of the Association for Computational Linguistics (NAACL 2018); New Orleans, LA2018.
- Wang H, Liu X, Xiao Y, Xu M, Xing EP. Multiplex Confounding Factor Correction for Genomic Association Mapping with Squared Sparse Linear Mixed Model. Methods. 2018;145:33-40. Epub 2018/05/01. doi: 10.1016/j.ymeth.2018.04.020. PubMed PMID: 29705210; PMCID: PMC6108326.
- Xie P, Wu W, Zhu Y, Xing EP. Orthogonality-Promoting Distance Metric Learning: Convex Relaxation and Theoretical Analysis. The 35th International Conference on Machine Learning (ICML 2018)2018.
- Xie P, Xing EP. A neural architecture for automated icd coding. InProceedings of the 56th Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers) 2018 Jul (pp. 1066-1076).
- Dan C, Leqi L, Aragam B, Ravikumar PK, Xing EP. The sample complexity of semi-supervised learning with nonparametric mixture models. InAdvances in Neural Information Processing Systems 2018 (pp. 9321-9332).
- Xie P, Zhang H, Zhu Y, Xing EP. Nonoverlap-promoting variable selection. InInternational Conference on Machine Learning 2018 Jul 3 (pp. 5413-5422).