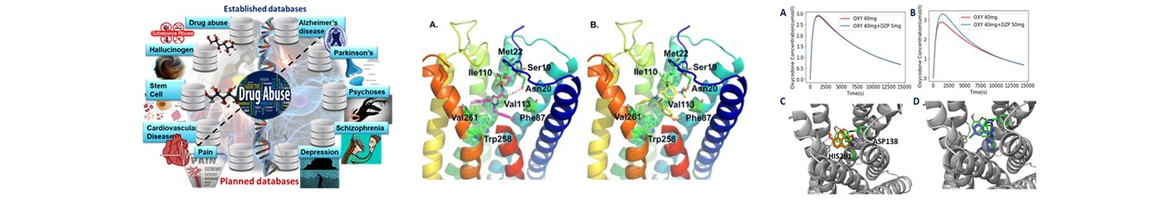
- Chen S, Feng Z, Wang Y, Ma S, Hu Z, Yang P, Chai Y, Xie, X.-Q*. Discovery of Novel Ligands for TNF-α and TNF Receptor-1 through Structure-Based Virtual Screening and Biological Assay. J Chem Inf Model. 2017 Apr 19. PMID: 28422491
- Bian Y, Feng Z, Yang P, Xie, X.-Q*. Integrated In Silico Fragment-Based Drug Design: Case Study with Allosteric Modulators on Metabotropic Glutamate Receptor 5. AAPS J. 2017 Jul;19(4):1235-1248.
- Cha-Molstad H, Yu JE, Feng Z, Lee SH, Kim JG, Yang P, Han B, Sung KW, Yoo YD, Hwang J, McGuire T, Shim SM, Song HD, Ganipisetti S, Wang N, Jang JM, Lee MJ, Kim SJ, Lee KH, Hong JT, Ciechanover A, Mook-Jung I, Kim KP, Xie, X.-Q*, Kwon YT*, Kim BY*. p62/SQSTM1/Sequestosome-1 is an N-recognin of the N-end rule pathway which modulates autophagosome biogenesis. Nature Communications 8, Article number: 102 (2017).
- Wang N, Wang L, Xie X.-Q. ProSelection: A Novel Algorithm to Select Proper Protein Structure Subsets for in Silico Target Identification and Drug Discovery Research. J Chem Inf Model. 2017 Nov 27;57(11):2686-2698. doi: 10.1021/acs.jcim.7b00277. Epub 2017 Oct 26. PMID:29016123
- Sauerwald N, Zhang S, Kingsford C, Bahar I. Chromosomal dynamics predicted by an elastic network model explains genome-wide accessibility and long-range couplings. Nucleic acids research. 2017 Apr 20;45(7):3663-73. PubMed PMID: 28334818; PubMed Central PMCID: PMC5397156.
- Cheng MH*, Torres-Salazar D*, Gonzalez-Suarez AD, Amara SG, Bahar I. (2017) Substrate transport and anion permeation proceed through distinct pathways in glutamate transporters.eLife 6: e25850.PMID: 28569666. (* equally contributed)
- Cheng MH, Garcia-Olivares J, Wasserman S, DiPietro J, Bahar I. (2017) Allosteric modulation of human dopamine transporter activity under conditions promoting its dimerization. J. Biol. Chem; in press.
- Lee J. Y., Feng Z., Xie X.-Q. , and Bahar I. (2017) Allosteric modulation of intact γ-secretase collective dynamics. Biophys. J. in revision.
- Li H, Sharma N, General IJ, Schreiber G, Bahar I. (2017) Dynamic Modulation of Binding Affinity as a Mechanism for Regulating Interferon Signaling. J Mol Biol; in press PMID:28648616.
- Li H, Chang Y-Y, Lee JY, Bahar I and Yang L-W. (2017) DynOmics: Dynamics of Structural Proteome and Beyond. Nucleic Acids Res. 45: W374-W380; PMID: 28472330.
- Ma S, Cheng MH, Guthrie DA, Newman AH, Bahar I, Sorkin A. (2017) Targeting of dopamine transporter to filopodia requires an outward-facing conformation of the transporter. Sci Rep. 7: 5399; PMID:28710426.
- Cheng, MH*, Garcia-Olivares, J*, Wasserman, S, DiPietro, J & Bahar, I. (2017) Allosteric modulation of human dopamine transporter activity under conditions promoting its dimerization. J. Biol. Chem 292:12471-12482; PMID: 28584050.
- Gur M, Cheng MH, Zomot E, Bahar I. (2017) Effect of dimerization on the dynamics of neurotransmitter: sodium symporters. J. Phys. Chem. B 121: 3657–3666. PMID: 28118712.
- Wang H, Aragam B, Xing EP. Variable Selection in Heterogeneous Datasets: A Truncated-rank Sparse Linear Mixed Model with Applications to Genome-wide Association Studies. KDD : proceedings. International Conference on Knowledge Discovery & Data Mining. bioRxiv. 2017 Jan 1:228106.
- Xu, X. Chai, H. Muthakana, X. Liang, G. Yang, T. Zeev-Ben-Mordehai and E. P. Xing, “Deep learning based subdivision approach for large scale macromolecules structure recovery from electron cryo tomograms.” The Twenty-fifth International Conference on Intelligence Systems for Molecular Biology (ISMB 2017). Bioinformatics (2017) in press.
- Zhou Z, Luo A, Shrivastava I, He M, Huang Y, Bahar I, Liu Z, Wan Y. Regulation of XIAP turnover reveals a role for USP11 in promotion of tumorigenesis. EBioMedicine. 2017 Feb 1;15:48-61.PubMed PMID: 28040451; PubMed Central PMCID: PMC5233825.
- Xie X-Q, Wang L, Wang J, Xie Z, Yang P, Ouyang Q. Neuropathology of Drug Addictions and Substance Misuse. San Diego, CA. Elsevier Inc.;. Chapter 19, In silico chemogenomics knowledgebase and computational system neuropharmacology approach for cannabinoid drug research.
- Ponzoni L, Zhang S, Cheng MH, Bahar I. Shared Dynamics of LeuT Superfamily Members and Allosteric Differentiation by Structural Irregularities and Multimerization. Phil Trans R Soc B. 2018;373. doi: 10.1098/rstb.2017.0177. PubMed PMID: 29735731; PMCID: PMC5941172.
- Al-Shedivat M, Wilson AG, Saatchi Y, Hu Z, Xing EP. Learning Scalable Deep Kernels with Recurrent Structure. Journal of Machine Learning Research. 2017;18:1-37. PMID: 30662374 PMCID: PMC6334642
Wenzel SE, Tyurina YY, Zhao J, Croix CMS, Dar HH, Mao G, Tyurin VA, Anthonymuthu TS, Kapralov AA, Amoscato AA, Mikulska-Ruminska K, Shrivastava IH, Kenny EM, Yang Q, Rosenbaum JC, Sparvero LJ, Emlet DR, Wen X, Minami Y, Qu F, Watkins SC, Holman TR, VanDemark AP, Kellum JA, Bahar I, Bayir H, Kagan VE. PEBP1 Wardens Ferroptosis by Enabling Lipoxygenase Generation of Lipid Death Signals. Cell. 2017;171(3):628-41. e26; PMCID: PMC5683852
Neiswanger W, Xing EP. Post-Inference Prior Swapping. The 34th International Conference on Machine Learning (ICML 2017); Sydney, Australia2017.
- Qin L, Zhang Z, Zhao H, Hu Z, Xing EP. Adversarial Connective-Exploiting Networks for Implicit Discourse Relation Classification. The 55th Annual Meeting of the Association for Computational Linguistics (ACL 2017); Vancouver, Canada.2017.
- Sachan M, Dubey A, Xing EP. From Textbooks to Knowledge: A Case Study in Harvesting Axiomatic Knowledge from Textbooks to Solve Geometry Problems. Proceedings of the 2017 Conference on Empirical Methods in Natural Language Processing (EMNLP 2017); Copenhagen, Denmark2017.
- Chang X, Yu Y, Yang Y, Xing EP. Semantic Pooling for Complex Event Analysis in Untrimmed Videos. IEEE Trans Pattern Anal Mach Intell. 2017;39(8):1617-32. Epub 2017/01/24. doi: 10.1109/tpami.2016.2608901. PubMed PMID: 28113653; PMCID: PMC5570670.
- Xie P, Deng Y, Zhou Y, Kumar A, Yu Y, Zou J, Xing EP. Analyzable Diversity-Promoting Latent Space Models. The 34th International Conference on Machine Learning (ICML 2017); Sydney, Australia2017
- Xie P, Singh A, Xing EP. Uncorrelation and Evenness: A New Diversity-Promoting Regularizer. The 34th International Conference on Machine Learning (ICML 2017); Sydney, Australia2017.
Xie P, Xing EP. A Constituent-Centric Neural Architecture for Reading Comprehension. The 55th Annual Meeting of the Association for Computational Linguistics (ACL 2017); Vancouver, Canada 2017.
- Yen IE, Huang X, Dai W, Ravikumar P, Dhillon I, Xing EP. Pd-Sparse : A Primal and Dual Sparse Approach to Extreme Multiclass and Multilabel Classification. The 23rd SIGKDD Conference on Knowledge Discovery and Data Mining (KDD2017); Halifax, Nova Scotia, Canada 2017.
- Zhang H, Zheng Z, Dai W, Q. H, Xing EP. Poseidon: An Efficient Communication Interface for Distributed Deep Learning on GPU Clusters. USENIX Annual Technical Conference; Santa Clara, CA, USA2017.
- Zhang K, Liu C, Zhang J, Xiong H, Xing EP, Ye J. Finding Structures of Large Matrices through Compression. The 23rd SIGKDD Conference on Knowledge Discovery and Data Mining (KDD2017); Halifax, Nova Scotia, Canada2017.
- Zhou Y, Yuan K, Yu Y, Ni X, Xie P, Xing EP, Xu S. Inference of Multiple-Wave Population Admixture by Modeling Decay of Linkage Disequilibrium with Polynomial Functions. Heredity (Edinb). 2017;118(5):503-10. Epub 2017/02/16. doi: 10.1038/hdy.2017.5. PubMed PMID: 28198814; PMCID: PMC5564381
- Lee S, Wang H, Xing EP. Backward genotype-transcript-phenotype association mapping. Methods. 2017 Oct 1;129:18-23. PMID: 28917724 DOI: 10.1016/j.ymeth.2017.09.004
- Xie P, Deng Y, Zhou Y, Kumar A, Yu Y, Zou J, Xing EP. Learning latent space models with angular constraints. InProceedings of the 34th International Conference on Machine Learning-Volume 70 2017 Aug 6 (pp. 3799-3810). JMLR. org
- Xie P, Salakhutdinov R, Mou L, Xing EP. Deep determinantal point process for large-scale multi-label classification. InProceedings of the IEEE International Conference on Computer Vision 2017 (pp. 473-482).
- Xie P, Poczos B, Xing EP. Near-Orthogonality Regularization in Kernel Methods. InUAI 2017 (Vol. 3, p. 6).
- Lee S, Görnitz N, Xing EP, Heckerman D, Lippert C. Ensembles of lasso screening rules. IEEE transactions on pattern analysis and machine intelligence. 2017 Nov 24;40(12):2841-52. PMID: 29989981